Thank you for visiting nature.com. You are using a browser version with limited support for CSS. To obtain the best experience, we recommend you use a more up to date browser (or turn off compatibility mode in Internet Explorer). In the meantime, to ensure continued support, we are displaying the site without styles and JavaScript.
Advertisement
Scientific Reports volume 14, Article number: 24799 (2024)
1233
Metrics details
This study aimed to analyze the molecular characteristics of insulin-like growth factor 1 (IGF1) gene in the testes of Tibetan sheep and its role in the testosterone synthesis and cell development. First, we cloned IGF1 gene for bioinformatics analysis, and the primary Leydig cells (LCs) of Tibetan sheep were isolated to explore its effect on the proliferation, apoptosis and function of LCs. Finally, the specific regulatory mechanism of IGF1 on LCs was analyzed by transcriptome sequencing. Results showed that overexpression of IGF1 increased the proliferation rate and decreased apoptosis of LCs. In addition, overexpression of IGF1 altered expression of genes related to testosterone synthesis and transformation and significantly increased amount of the final product testosterone. Mechanistically, IGF1 stimulated the expression of the proliferating cell nuclear antigen and IGF1R and promoted the proliferation of LCs via the PI3K/Akt signaling pathway. Collectively, what should be clear from the results reported here is that IGF1 might play roles in the proliferation or differentiation and testosterone synthesis of LCs. These findings add to our understanding on the regulation of testosterone synthesis in sheep and other mammals.
Sheep industry occupies an important position in China’s livestock industry, and its production level and breeding efficiency are closely related to reproductive traits1. As the most valuable germplasm resource distributed in high altitude areas, Tibetan sheep plays an important role in the economic income of local farmers and herdsmen and the function of grassland ecosystem. However, due to its low fecundity and late sexual maturity2,3, it is urgent to decipher its reproductive physiological characteristics and reproductive-related problems. Leydig cells, as the most important and distributed cell group in the Leydig region, play an important role in sperm formation, maturation, motility, and male fertility maintenance. It can synthesize and secrete a large of steroid hormones4,5, providing a suitable environment and nutrition support for the development of male germ cells. For example, androstenedione plays an important regulatory role in spermatogenesis and sperm maturation and promotes sperm formation and sperm cell differentiation to prevent stagnation of sperm development6. In the absence of testosterone stimulation, spermatogenesis does not proceed beyond meiosis stage7. Additionally, in the sexually mature mammalian testis, androgen can significantly up-regulate the expression of genes related to the establishment and function maintenance of the blood-testis barrier in Sertoli cells, thus providing an immunosuppressive microenvironment for meiosis and subsequent normal development of spermatogenic cells8. Therefore, it is highly significant to investigate the mechanism behind the proliferation, apoptosis, and maturation of LCs.
Insulin-like Growth Factor 1 (IGF1) is a significant member of the Insulin-like growth factors (IGFs) family, was extensively studied because its crucial role in sex determination, gonadal development, and reproduction in mammals9. In recent years, with the development of molecular biology, IGF1 and IGF2 have been identified, which involved in protein synthesis, nutrient metabolism, but also in cell growth, differentiation, apoptosis, and embryonic development, highlighting their importance. It is worth noting that IGF1 is a polypeptide hormone composed of 70 amino acid residues. It mainly acts on the reproductive system and immune system, synthesized by the liver and other tissues, and regulated by growth hormone (GH). Previous studies have demonstrated that IGF1 expressed in various types of testicular cells10, including germ cells, Leydig cells, Sertoli cells, and promoted the proliferation of various cell types, such as smooth muscle cells11, fibroblast cells12 and germ cells13. Former researchers have found that IGF1, together with GDNF, FGF2 and bFGF, can support the maintenance and proliferation of spermatogonia stem cells14,15, and that IGF1 secreted by mouse testicular interstitial progenitor cells promotes the proliferation of immature Sertoli cells through the IGF1/PTEN/PI3K signaling pathway in a paracrine manner, thereby regulating testicular size and sperm production16. Nonetheless, studies on IGF1 have only focused on goat, pig, mice, and humans, less attention has been placed on IGF1 on Tibetan sheep testes.
Therefore, in our study, the IGF1 gene was cloned and sequence analysis was performed on it. Subsequently, cell transfection experiments were designed to determine the impacts of IGF1 gene on the proliferation and apoptosis of LCs and the expression of testosterone synthesis related genes, and testosterone concentration of Tibetan sheep testes. This not only offers valuable evidence for the role of IGF1 in the testosterone synthesis of Tibetan sheep but also provides useful clues for the molecular mechanism of regulating the proliferation and apoptosis of LCs in Tibetan sheep.
Protocols of this study were approved and consented on by the Animal Committee of Gansu Agricultural University (Ethical approval No GSAU-Eth- ASF2022-008). All animals received care in accordance with the recommendations of the National Institutes of Health Guide for Care and Use of Laboratory Animals and adhered to the ARRIVE guidelines.
Primer 6.0 software (Premier Biosoft International, Palo Alto, CA, USA) was used to design primers according to the predicted mRNA sequence of sheep IGF1 gene (GenBank Accession: No. NM_001009774.3). Their specificity was evaluated using the basic local alignment search tool (BLAST) online program (NCBI, U.S. Library of Medicine, Bethesda, MD, USA; http://blast.Ncbi.nlm.nih.gov/Blast.cgi). The cDNA derived from Tibetan sheep testis samples was used as the template. The PCR reaction system (20 µL) included 1.5 µL of cDNA, 0.5 µL each of forward and reverse primers, 10 µL of high-fidelity DNA polymerase (1U/mL), and 7.5 µL of dd H2O. The reaction consisted of 1 cycle of 94 °C for 5 min, followed by 32 cycles of 94 °C for 30 s, 53 °C for 30 s, and 72 °C for 2 min. Separation of the amplified products was achieved using a 1% agarose gel. The target fragment was purified and recovered through an agarose gel DNA recovery kit (TIANGEN, Beijing, China). Subsequently, the fragment was ligated with the pEASY-Blunt vector and transformed into Trans1-T1 competent cells (TransGen Bioiech, Beijing, China). The positive clones were selected on LB-ampicillin plates, and identified by PCR using a 2 × EasyTaq PCR SuperMix (TransGen, Beijing, China). Eight independent positive clones were arbitrarily selected and sequenced by the Tsingke Biotechnology Co. Ltd. (Xi’an, China).
To search for homologous sequences in NCBI, the BLAST method was employed. By utilizing DNAMAN 9.0 software, multiple alignments of the IGF1 CDS sequence from various species were conducted. For the analysis of the entire ORF sequence of the IGF1 gene, the ORF Finder online software provided by NCBI database was utilized. To construct the phylogenetic tree, the neighbor-joining method in MEGA11.0 software was employed. Several online software tools were also utilized for further analysis. Prot Param (accessible at http://web.expasy.org/protparam/) was used to predict and analyze basic physicochemical properties of IGF1. The Signal-4.1 software (available at https://services.healthtech.dtu.dk/service.php?SignalP-4.1) aided in analyzing the signal peptide. SOPMA (found at https://npsa-prabi.ibcp.fr/cgi-bin/npsa_automat.pl?page=npsa_sopma.html) was utilized to examine the secondary protein structure. Lastly, Phyre2 (accessible at https://bioinf.cs.ucl.ac.uk/psipred/) was used to predict the tertiary protein structure of IGF1.
Three one-month-old healthy male Tibetan sheep with the same feeding conditions were selected as experimental individuals (Xiahe, Gansu, China; altitude > 3,000 m; latitude 34°32′ N, longitude 101°54′ E). These sheep were brought back to the laboratory and were intravenously injected with pentobarbital sodium to no heartbeat, no blink reflex, then castrated. After castration, the testicles were collected and immersed in Hanks’ balanced salt solution (HBSS) containing 100 IU mL of penicillin-streptomycin at a temperature of 4 ℃. The surface envelope was then incised, ensuring aseptic conditions by using scissors to remove any contaminated portions. Tissue samples were obtained from the central region of testicle sample in a 50 mL centrifuge tube containing 20 mL of collagenase type IV (0.25 mg/mL) and 1 mL of DNase I (7 mg/mL). Digestion was terminated by adding DMEM/F12 medium (Gibco, New York, USA) supplemented with serum. Single testicular cells were obtained by passing the suspension through a 70 μm cell strainer. The cells were then cultured in DMEM/F12 medium supplemented with 15% fetal bovine serum (Gibco, New York, USA) with 1% penicillin-streptomycin solution (Gibco, New York, USA). The adherent and proliferating cells were retained, and then the cells were purified by percoll method17and differential adhesion method18. Briefly, after the cells were adhered and differentiated, they were digested with 0.25% trypsin, and the digestion was stopped by adding an equal volume of fetal bovine serum to collect the cell suspension. Next, the cell suspension was added to the Percoll gradient separation solution with varying volume fractions, and centrifuged at 3000 rpm for 30 min. Then, the cell solution between the 34–60% gradient was transferred to a new centrifuge tube. An appropriate amount of PBS containing 1% penicillin-streptomycin solution was added, and the tube was centrifuged at 1000 rpm for 15 min. The supernatant was discarded to obtain relatively pure LCs. The volume fractions of percoll cell separation solution (Solarbio, Beijing, China) were 21%, 26%, 34% and 60%, respectively.
The IGF1- siRNA interference sequence was designed and synthesized based on the cloned sequence of Tibetan sheep IGF1. Shanghai Genepharma Biological Co., Ltd. (Table 1) was responsible for this task, while the construction of the overexpression vector (pcDNA3.1 (+)-IGF1) was completed by Suzhou Genewiz Biological Co., Ltd. The plasmid mini-extraction kit (Tiangen, Beijing, China) was utilized to extract the plasmid, following the provided instructions. Double-enzyme digestion with NheI and NotI restriction enzymes was then performed, and the digestion products were evaluated using 2% agarose gel electrophoresis. Transfection commenced when the LCs reached a confluence level of 60–70%. The Lipofectamine 3000 kit instructions (Invitrogen, California, USA) were strictly followed during the transfection procedure.
After isolation, purification, and characterization, only the LCs of F4 generation were selected for transfection once they achieved a confluence level of 60–70%. Initially, 2 µL of IGF1 overexpression plasmid DNA at a concentration of 1000 ng/µL and 5 µL of the transfection reagent Lipofectamine 3000 (Invitrogen, California, USA) were individually diluted with 250 µL of serum-free medium opti-MEM (Gibco, New York, USA). The two solutions were gently mixed and incubated for 5 min at room temperature. Subsequently, the two liquids were combined and incubated at room temperature for 20 min, resulting in the formation of a DNA-Lipofec-Tamine 3000 complex. The cells were then washed twice with preheated PBS before adding 500 µL of the complex to a six-well plate containing 1.5 mL of pre-added opti-MEM. To ensure thorough mixing, the six-well plate was lightly shaken. The plate was then placed in a 37 °C, 5% CO2 incubator for 6 h, followed by replacement with DMEM/F12 medium supplemented with 15% FBS for culture. It’s important to note that the siRNA dosage for transfection was 150 nM/well, and the lipofectamine 3000 dosage was 5 µL/well. The remaining steps were like those involved in plasmid transfection.
To determine the viability of cells, the CCK-8 reagent obtained from MedChen Express based in Shanghai, China, was utilized. The LCs in the logarithmic phase were carefully selected and subjected to trypsin digestion, followed by cell counting. Subsequently, the cells were seeded into a 96-well plate with a density of 1 × 103 cells per well, thereafter allowing them to incubate for 24 h. Following the initial incubation, transfection was performed for durations of 24, 48, and 72 h. After the transfection periods, each well received an addition of 10 µL of the CCK-8 reagent and incubated for one hour at the temperature of 37 °C Eventually, the optical density (OD) value of each well was measured at 450 nm through the employment of enzyme-linked immunosorbent assay (ELISA).
We employed flow cytometry to assess labeled necrotic and apoptotic cells using Annexin V-FITC/PI (Q2 + Q3). Briefly, we generated a single cell suspension by treating cells with EDTA-free trypsin and subsequently rinsed them with pre-chilled PBS. Afterwards, the cells were incubated with 300 µL of binding buffer at a density of approximately 5 × 105 cells per tube. Each tube received 5 µL of Annexin V-FITC, which was thoroughly mixed. Then, 5 µL of PI solution was added, mixed thoroughly, and shielded from light for 10 min. Finally, flow cytometry was employed to analyze the liquid in the tube.
Following the provided instructions, after transfection of LCs, they were homogenized and lysed using a radioimmunoprecipitation assay (RIPA) protein extraction kit (Solarbio, Beijing, China). The concentration of the proteins was determined using a commercial bicinchoninic acid (BCA) protein assay (Solarbio, Beijing, China). Subsequently, the extracted proteins underwent denaturation with 5x SDS sample buffer (Solarbio, Beijing, China). To separate the proteins, a 12% sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) technique was employed, and they were subsequently transferred onto polyvinylidene difluoride (PVDF) blotting membranes (Beyotime, Shanghai, China). The membranes were sealed by using phosphate buffered saline tween-20 (PBST) with 5% non-fat milk for 2 h at room temperature. Antibody specific information is displayed in Supplementary Table S1. Following the PBST wash, the membranes underwent incubation with goat anti-rabbit IgG/HRP antibody for 2 h at 37 °C. Visualization of the bands was accomplished using an enhanced chemiluminescence (ECL) kit (NCM Biotech, Suzhou, China) after PBST washing. The gray value of the band obtained after western blot analysis was scanned and determined using AlphaEaseFC 6.0 image analysis software. The relative expression of the target protein was the ratio of the gray value of the target protein to the gray value of β-actin.
In brief, aspirate 2 mL of Leydig cell suspension and transfer it into a 6-well culture plate. After a pre-culture period of 24 h, discard the culture medium. Wash the cells with PBS, then permeabilize the cell membrane using 0.3% Triton-X 100, allowing it to stand at room temperature for 10 min. Following this, block the cells with 5% bovine serum albumin (BSA; Solarbio, Beijing, China) and incubate the plate overnight at 4 °C with a polyclonal rabbit primary antibody against HSD3B1 (1:200 dilution; Bioss, Beijing, China). Subsequently, incubate the plate with CY3-labeled goat anti-rabbit IgG (1:400; Servicebio, Wuhan, China) for 1 h at room temperature in the dark. Wash the plate three times with PBS and treat it with 4’,6-diamidino-2-phenylindole (DAPI; Servicebio, Wuhan, China) for 10 min in the dark. After sealing, place the slides under a fluorescence microscope (Nikon, Eclipse C1, Tokyo, Japan) for observation. Experiments were conducted with four biological replicates, each comprising two technical replicates. Digital images were captured using CaseViewer software (3DHISTECH, Budapest, Hungary).
Supernatants and cells transfected for 48 h were harvested and testosterone concentration was determined by a commercial enzyme-linked immunosorbent assay (ELISA) kit (JM-07849S2, Jiangsu, China) following the manufacturer’s recommendations. Optical density was measured at 450 nm by a microplate reader (BioTek, Winooski, VT, USA). The intra-assay and inter-assay coefficients for androgen determination in this laboratory were less than 10 and 5%, respectively.
The total RNA of LCs after transfection were extracted using TransZol (TransGen Biotech, Beijing, China) according to manufacturer’s instructions. The integrity of RNA was determined by 1% agarose gel electrophoresis, whereas the concentrations and quality of RNA samples were examined by NanoDrop 2000 (Thermo Fisher Scientific, Waltham, MA, USA)) and Agilent 2100 (Agilent, Santa Clara, CA, USA), respectively. Next, Evo M-MLVRT Kit with gDNA Clean for RT-qPCR (Accurate Biotech, Hunan, China) was used to reverse transcribe the RNA into cDNA, and SYBR Green Premix Pro Taq HS qPCR Kit (Accurate Biotech, Hunan, China) was used to perform qPCR on the Roche LightCycler96 in accordance with the manufacturer’s protocols. The relative mRNA expression of each gene was normalized to β-actin (a housekeeping gene) mRNA and calculated using the 2–ΔΔCt method19. The primer pairs of selected genes were listed in Supplementary Table S2.
RNA extraction and cDNA library preparation LCs were gathered 48 h post transfection, followed by two PBS washes. Subsequently, each well underwent treatment with 1 mL Trizol on ice, leading to repetitive blowing, and thereby resulting in the extraction of total RNA after achieving complete lysis. The acquired total RNA samples were assessed for purity and concentration using the Nano Drop spectrophotometer and Agilent 2100 bioanalyzer, respectively. Moreover, a 1.5% agarose gel electrophoresis was conducted to evaluate the integrity and ratio of the 28 S and 18 S bands. Concurrently, RT-qPCR was utilized to measure the efficiency of overexpression and interference. Next, eight total RNA samples (four from each group) were selected to create cDNA library preparation. The construction of transcriptome libraries was achieved using the VAHTS Universal V5 RNA-seq Library Prep Kit. After library preparation, the Illumina Novaseq 6000 sequencing platform was employed to sequence the library, generating 150 bp paired-end reads. The initial processing of the raw fastq format reads involved the use of fastp to eliminate low-quality reads, establishing clean reads for subsequent analyses (Chen et al., 2018). Upon obtaining the clean reads, they were aligned to the reference genome of sheep (Oar_v1.0, GCF_002742125.1) using HISAT220. Following this, FPKM values were computed for each specific gene, and read counts were determined utilizing HTSeq-count21.
Differential expression analysis was conducted using DESeq222. The threshold for significantly differentially expressed genes (DEGs) was set as a false discovery rate (FDR) < 0.05 and a |log2FC| ≥ 1.0. R (v 3.2.0) was used to perform hierarchical cluster analysis of DEGs to visually display the expression patterns of genes in different samples. Moreover, GO (Gene Ontology) and KEGG (Kyoto Encyclopedia of Genes and Genomes) pathway enrichment analyses of DEGs were performed to screen the significant enriched term using R (v3.2.0), respectively23. In addition to GO and KEGG analyses, GSEA (Gene Set Enrichment Analysis) software was utilized to conduct gene set enrichment analysis.
All statistical analyses were performed by one-way analysis of variance (ANOVA) using SPSS 21.0 software and all data were expressed as mean ± standard error (Mean ± SE). P < 0.05 and P < 0.01 indicating statistically significant and extremely significant, respectively.
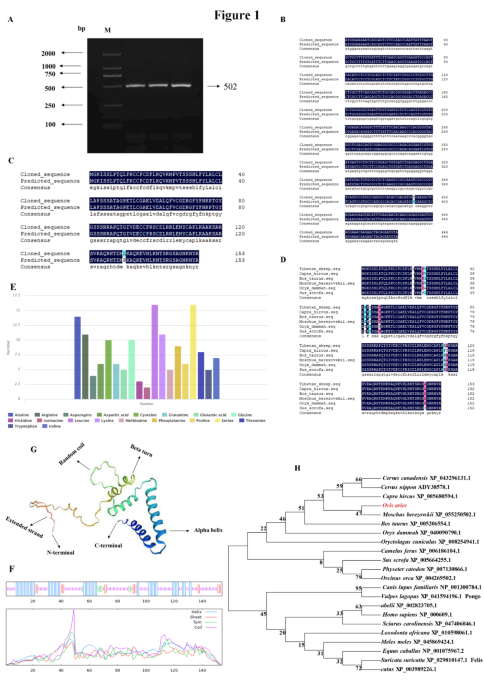
The target fragment of size 502 bp was obtained by the RT-PCR method using Tibetan sheep testes cDNA as templates (Fig. 1A). Through the ORF Finder online software analysis in the NCBI database, it was found that the cloned cDNA sequence included a 465 bp open reading frame (ORF) translatable to 154 amino acids, with an ATG (M) start codon and a TAG stop codon (Supplementary Fig. S1A). The resulting full-length CDS sequence of Tibetan sheep IGF1 has been deposited in Gen Bank (accession no. OR790200). The CDS sequence alignment showed that the cloned Tibetan sheep IGF1 sequence had one base substitution (C→T) at nucleotide position 326 compared with the predicted sheep IGF1 sequence from the National Centre for Biotechnology Information (NCBI) database (GenBank no. NM_001009774.3; Fig. 1B). The homology comparison results of amino acid sequences showed that Proline (Pro) changed into Leucine (Val) in Tibetan sheep IGF1 sequence compared with sheep sequence (NP_001009774.1; Fig. 1C). The results of amino acid sequence alignment are shown in Fig. 1D.
At the nucleotide level (Supplementary Fig. S1B), IGF1 cDNA sequence was composed of 22.58% base A (105), 21.71% base T (101), 26.45% base C (123), and 29.26% base G (136). The amino acid composition revealed that Leucine is the majority amino acid (10.40%), followed by Leucine (9.71%), Alanine (9.10%), and Arginine (7.10%) (Fig. 1E). The predicted physicochemical properties revealed that the molecular formula of IGF1 protein was C741H1184N214O216S15, with a molecular weight of 17027.76 u, a theoretical isoelectric point of 9.36 and an instability coefficient of 52.85, which is an unstable protein. The total average hydrophilicity was − 0.19, the lipid coefficient was 67.86, and the number of positively and negatively charged residues was 22 and 11, respectively. The Tibetan sheep IGF1 protein does not have a predicted transmembrane region, but there is a 65.53% probability of the presence of a predicted signal peptide with type SP (Sec / SPI) and the sequence being MGKISSLPTQLFKCCFCDFLKQVKMPVTSSSHLFYLALCLLAFSSSATA. The cutting site is located at 49–50 with a probability of 50.16% (Supplementary Fig. S1C-D).
Cloning results of Tibetan sheep IGF1 gene. (A) PCR amplification products of the Tibetan sheep IGF1 coding sequence (CDS). M, DL2000 marker; 1–2, PCR product. (B) Alignment of NCBI nucleotide sequence and cloned nucleotide sequence of Tibetan sheep IGF1. (C) Amino acid composition. (D) Multiple alignment of IGF1 amino acid sequences from 6 different mammalian species. (E) The amino acid composition. (F) Secondary molecular structure. Different lines with different colors denote different secondary structures: blue, alpha helix; red, extended strand; green, beta turn; purple, random coil. (G) Tertiary molecular. (H) Neighbor-joining phylogenetic tree based on nucleotide sequences of IGF1 gene among different mammals. Original blots are presented in Supplementary Fig. S2.
Secondary structure predictions for IGF1 protein showed that it was a protein with mixed secondary structures, including 33.77% α-helix, 51.96% random coil, 4.55% β-turn and 9.74% extended strand (Fig. 1F, G). The tertiary molecular structure analysis of the coded protein indicated that it was similar with secondary structure, consisting of α-helix, β-turn, random coil, and extended strand (Supplementary Fig. S1E). The phylogenetic tree was constructed based on nucleotide sequences of IGF1 from Tibetan sheep and other known mammals, the tree showed that Tibetan sheep IGF1 sequence was the most closely related to Moschus Berezowski (Fig. 1H).
The primary LCs of Tibetan sheep were isolated and purified, and the morphology was shown in Fig. 2A. Here, we found that purified LCs could express LCs marker genes HSD3B1 (Fig. 2B). Immunofluorescence staining of LCs with specific antibody HSD3B1 antibody showed that purified LCs could synthesize and express LCs-specific binding protein HSD3B1 (Fig. 2C). The results indicate that the cell purity exceeds 80% and can be used for subsequent experiments and analysis. The result of double digestion of pcDNA3.1 (+)-IGF1 was electrophoresed on a 2% agarose gel, as shown in Fig. 2D. The size of the digestion product was consistent with the theoretical value.
Identification of LCs, evaluation of recombinant plasmid and detection of transfection efficiency. (A) The primary Leydig cells of Tibetan sheep. (B) PCR results of HSD3B1 and GAPDH gene. (C) Immunofluorescence staining of primary LCs of Tibetan sheep (200 μm). (D) Evaluation of recombination plasmids. (E–G) Detection of transfection efficiency. Original blots are presented in Supplementary Fig. S2.
The LCs transfected for 24, 48 and 72 h were harvested, and the expression level of IGF1 mRNA in the transfected Tibetan sheep LCs was detected by RT-qPCR. The results showed that compared with the control group (NC), the expression of IGF1 was downregulated in the si-IGF1-1 and si-IGF1-2 transfected LCs, but the silencing effect was most obvious after transfection for 48 h (Fig. 2G). Comparing the transfection efficiencies of the two siRNAs transfected for 48 h, si-IGF1-1 has the best silencing effect (Fig. 2E). Compared with the empty vector group, the expression of IGF1 in pcDNA3.1(+)-IGF1 of the transfection group was upregulated after transfection for different periods (Fig. 2F). However, the upregulation was most obvious at 48 h, indicating that the over-expression efficiency of transfection at 48 h was the best. Considered together, these data indicate that pcDNA3.1(+)-IGF1 and si-IGF1 could effectively promote and inhibit the expression of IGF1 in LCs at the transcriptional level, respectively.
CCK-8 assay showed that compared with the empty vector group, there was no significant difference in cell viability in the pcDNA3.1(+)-IGF1 group after 24 h of transfection, while cell viability was significantly increased after 48 and 72 h of transfection (P < 0.05). The proliferation of LCs obviously reduced by the si-IGF1 group at 48 and 72 h (P < 0.05) (Fig. 3A). Flow cytometry was used to determine the changes in cell apoptosis after 48 h of transfection, and results showed that the apoptosis rate of the pcDNA3.1(+)-IGF1 group was significantly lower than that of the empty vector group (P < 0.05). The apoptosis rate of the si-IGF1 group was higher than that of the NC group (P < 0.05). The results of RT-qPCR demonstrated that the expression of BAX was significantly downregulated, whereas the expression of PCNA, BCL2 and CDK2 were significantly upregulated (P < 0.05), compared to the empty vector group (P < 0.05). After silencing of IGF1 gene, the results contrasted with the overexpression results (Fig. 3B-C). Collectively, these results suggest that overexpression of IGF1 promotes LCs proliferation and inhibits their apoptosis.
RT-qPCR and Western blot were employed to assess the expression of PI3K-Akt signaling pathway receptor IGF1R mRNA and protein after overexpression or silencing of IGF1 (Fig. 3D-E). These results showed that pcDNA3.1(+)-IGF1 significantly increased the expression of IGF1R mRNA and protein (P < 0.05) compared with the empty vector group and si-IGF1 remarkably decreased their expression (P < 0.05) compared to the control group (original blots are presented in Supplementary Fig. S3).
Effect of IGF1 gene silencing and overexpression on the proliferation, apoptosis, and cycle of LCs. (A) CCK-8 detects the cell proliferation rate after overexpression and silencing of IGF1. (B) Flow cytometry was used to detect the apoptosis of LCs after overexpression and silencing of IGF1. (C) The effect of overexpression and silencing of IGF1 gene on the expression of proliferation, cycle, and apoptosis related genes. (D, E) The expression of IGF1R mRNA and protein. Original blots are presented in Supplementary Fig. S3.
After 48 h of transfection, the expression levels of genes and proteins related to testosterone synthesis in LCs were detected by RT-qPCR and Western blot, respectively. These results showed that pcDNA3.1(+)-IGF1 group significantly enhanced STAR, CYP11A1, and HSD17B3 mRNA and protein (P < 0.05) compared with the empty vector group, whereas the expression of SRD5A2 mRNA and protein were significantly downregulated (P < 0.05), and si-IGF1 remarkably opposite their expression (P < 0.05) compared to the control group (Fig. 4B-D). The testosterone kit was then used to determine the content of testosterone in LCs after 48 h of transfection. Results revealed that the testosterone content in the supernatant of the pcDNA3.1 (+)-IGF1 group was significantly increased, but there was no significant difference in the intracellular testosterone content (P < 0.05). After silencing the IGF1 gene, the result was the opposite of overexpression (Fig. 4A).
The effect of IGF1 on testosterone content of Leydig cells. (A) Testosterone content in culture medium and intracellular. Graphic area represents testosterone content. (B, D) Changes in expression of HSD11B1, CYP11A1, HSD3B1, SRD5A2, HSD17B3, and STAR after overexpression or silencing of IGF1 gene. (C) Western blot analysis of SRD5A2, HSD17B3, CYP11A1 and β-actin expression in the four groups, original blots are presented in Supplementary Fig. S3.
In the transcriptomic analysis, 65,685,881 and 63,575,740 high quality clean reads were obtained from the LCs and pcDNA3.1(+)-IGF1, respectively. A total of 65,685,881 (LCs), and 63,575,740 (PcDNA3.1(+)-IGF1) clean reads were mapped to the ovine genome with mapping rations of 97.30% and 97.26%, respectively. Among the total mapped reads, 91.35%and 83.02% form the LCs and PcDNA3.1(+) -IGF1 were unique matches to the ovine genome (Supplementary Table S3). These results showed that the sequencing quality was high enough for the subsequent quantitative analysis. In total, 1,040 DEGs were identified in groups of pcDNA3.1(+) -IGF1vs LCs, including 492 up-regulated genes and 548 down-regulated genes (Fig. 5A, Supplementary Table S4). These DEGs were mostly related to Leydig cells growth, development, and function, namely, Proliferating Cell Nuclear Antigen (PCNA), Marker of proliferation Ki-67 (MKI67), Insulin-like Growth Factor 1 Receptor (IGF1R), Steroid 5 alpha-reductase 2 Gene (SRD5A2), Superoxide dismutase 3 Gene (SOD3) and Cytochrome P450 family 17 subfamily A member 1 Gene (CYP17A1). To further explore the dynamics expression these genes in the overexpression group, we performed cluster analysis. Here, we identified 7 clusters (Fig. 5B). The difference, that is, the genes with high expression in the NC group were gradually down-regulated in the pcDNA3.1(+) -IGF1 group, while the genes with low FPKM in the pcDNA3.1(+) -IGF1group were gradually up-regulated in the NC group. This result indicates that overexpression of IGF1 gene can inhibit the expression of number genes or induce the expression of another genes, and the changes of these genes may be to regulate the proliferation and differentiation of LCs. Correlation heat map revealed that the gene expression profiles in the overexpression group was differed significantly from the control group (Fig. 5C).
Differential Genes expression profiles. (A) Volcano plot showing DEGs expression in two groups. (B) Cluster analysis of DEGs in two groups. (C) Correlation heat map in two groups.
The GO classification results showed that 358 GO terms significantly enriched in the pcDNA3.1(+) -IGF1 group compared with the LCs group (P<0.05). Among them, most of DEGs were enriched in terms such as biological process, were annotated to cellular response processes and metabolic process; in cellular component, were annotated to extracellular matrix components, MHC protein complexes, and other processes; in molecular function, were annotated to processes such as chemokine activity, chemokine receptor binding, ATP-dependent activity (Supplementary Table S5) (Fig. 6A).
The GO and GSEA annotation of DEGs in the two groups. (A) GO enrichment terms. (B) KEGG enrichment terms. (C–F) GSEA enrichment result maps in two groups, respectively. BP: biological process; CC: cellular component; MF: molecular function.
The KEGG pathway analysis results showed that 40 pathways were significantly enriched in the DEGs of the overexpression group vs. LCs group comparison (P < 0.05) (Supplementary Table S6). KEGG enrichment analysis showed that DEGs in pcDNA3.1(+) -IGF1 vs. LCs group were mainly enriched in Aldosterone synthesis and secretion, PI3K-Akt signaling pathway, ABC transporters, and MAPK signaling pathway (Fig. 6B). In GSEA, we screened the significantly enriched functional pathways with the criteria of | NES |>1, P < 0.05, and FDR < 0.25. As shown in Fig. 6C-F, the PI3K-Akt signaling pathway and Cholesterol synthesis were significantly activated in the overexpression group, and the gene set of the overexpression group was up-regulated. The Apoptosis and the lipolysis metabolism pathway were significantly inhibited in the overexpression group, and the gene set of the overexpression group was down-regulated. We further analyzed the expression of vital genes in these pathways and found that the expressions of PCNA, MKI67, CYP11A1, HSD11B1, and HSD17B3 were significantly higher in the pcDNA3.1(+) -IGF1 group than in the LCs group (P < 0.05) (Supplementary Table S7). In conclusion, IGF1 gene may regulate testosterone levels by promoting the expression of genes related to Leydig cell proliferation and testosterone synthesis.
To further analyze the regulatory mechanism of IGF1 in testosterone production, we identified several gene sets associated with the process of testosterone synthesis by integrating the database-based and literature-based functional annotations, such as cholesterol synthesis, androgen synthesis and testosterone secretion. Among these, a total of 181 genes related to testosterone synthesis were detected in the overexpression groups (Supplementary Table S8).
Identification of testosterone synthesis related genes and validation of RNA-Seq by RT-qPCR analysis. (A) The PPI network of gene sets enriched on testosterone synthesis-related pathways. (B) The gene-pathway network according to cholesterol synthesis and androgen synthesis-related genes and their enriched pathways. (C) OE and NC represent pcDNA3.1(+) -IGF1 and LCs, respectively.
The PPI network of genes closely associated with testosterone synthesis, with 99 nodes and 455 edges, is presented in Fig. 7A. The top hub genes (degree ≥ 10) in the network were cholesterol transformation related genes (i.e., STAR, CYP11A1, HSD3B1, HSD17B3, and CYP11B1), cholesterol synthesis related genes (i.e., HMGCR, LDLR), genes related to gonadotropin secretion (GnRH1). Among them, most of genes were mainly involved in the synthesis and transformation of cholesterol, the synthesis of progesterone and the release of testosterone. Specifically, 8 genes were involved in aldosterone synthesis and secretion, 5 genes were involved in GnRH secretion, 12 genes were involved in MAPK signaling pathway, 4 genes were involved in Cortisol synthesis and secretion, 7 genes were involved in Signaling pathways regulating pluripotency of stem cells, 8 genes were involved in Metabolic pathways, and 2 genes were involved in Steroid hormone biosynthesis (Fig. 7B). To validate the transcriptomic data, 10 DEGs were randomly selected for RT-qPCR detection. As shown in Fig. 7C, the results showed that the expression patterns of the genes were identical to those in the RNA-seq data, indicating that the sequencing results were accurate and reliable.
As an important member of the Insulin-like growth factors (IGFs) family, IGF1 has been demonstrated to have a crucial role in sex determination, gonadal development, and reproduction in mammals24. Previous studies have indicated that the absence of IGF1 can result in significant alterations in the development of adrenal cortex and testis, leading to impaired testosterone synthesis and spermatogenesis25. In previous studies, we found that IGF1 was mainly localized in Leydig cells and Sertoli cells (SCs) of Tibetan sheep, and the expression level gradually decreased with age, showing the same trend as testosterone in serum3,10. Based on these findings, it can be postulated that IGF1 may play an indispensable role in testes development and testosterone synthesis in Tibetan sheep. Therefore, we cloned the full-length coding sequence (CDS) of the IGF1 gene using cDNA from Tibetan sheep testicular tissue as a template. A comparison of the CDS sequence of IGF1 with the sheep sequence from the National Centre for Biotechnology Information (NCBI) database revealed only one base substitution (C→T), suggesting that the IGF1 gene is highly conserved in the evolutionary process. Additionally, when comparing the amino acid sequences, it was observed that the IGF1 sequence of Tibetan sheep had a change from Proline (Pro) to Leucine (Leu), further supporting the high conservation of the IGF1 gene in the evolutionary process. Proline, known for its rigidity, can be incorporated to increase the rigidify of flexible regions to enhance the thermostability of proteins26. It is hypothesized that the substitution of Proline with Leucine may contribute to the enhancement of protein thermostability in Tibetan sheep.
Apoptosis is essential for the elimination of unnecessary or overproduced cells during mammalian spermatogenesis, which can be triggered by a variety of stimuli, including cytokines, hormones, viruses, and toxic substances27. B-cell lymphoma-2 (BCL2), an oncogene isolated from mouse lymphocyte, regulates various biological processes such as including many embryonic, adult tissues and immune defense by mediating apoptosis. In experiments using BCL-w protein mutant mice, it was discovered that the downregulation of BCL2 resulted in the death of SCs28. It is reported that BAX gene knockout, the apoptosis of testicular germ cells in neonatal mice is significantly inhibited, and mature haploid sperm could not be produced29. Additionally, one study also found that IGF1 can induce SCs apoptosis by increasing the expression of BAX and decreasing the expression of BCL230. In the present study, we found that pcDNA3.1(+)-IGF1 enhanced the BCL2 mRNA and decreased the BAX mRNA in LCs. Opposite results were observed when LCs were treated with si-IGF1. It is worth noting that related study found that after IGF1 gene was knocked out, the proapoptotic factor BAX was upregulated and the anti-apoptotic factor BCL2 was downregulated in sheep Sertoli cells, which was completely consistent with our results31. The above results imply that IGF1 promotes the proliferation of LCs via regulating the expression of BAX and BCL2.
Cell proliferation is primarily driven by altered cell cycle progression, with PCNA and CDK2 being important regulators in the cell cycle. PCNA, also known as proliferating cell nuclear antigen, is a core protein involved in DNA synthesis and repair, playing a crucial role in the regulation of the s-phase of germ cell meiosis32. Ma et al. further revealed that PCNA proteins form cyclic trimers around DNA during DNA replication, enhancing the continuity of DNA synthesis33. Mouse Tspyl5 promotes spermatogonia proliferation through enhancing PCNA-mediated DNA replication34. Silencing PCNA gene expression can inhibit the proliferation of thyroid cancer cells and promote their apoptosis35. During the meiosis of germ cells, CDK2 is a key regulatory protein that drives cells to enter the S phase through the G1/S phase checkpoint to complete DNA synthesis. Overexpression of CDK2 accelerates the transition from G1 to S phase, thereby promoting excessive cell proliferation36. Recently, a study revealed that IGF1 played a positive role in cell cycle progression, accompanied by enhanced mRNA expression of PCNA and CDK210. In the current study, it was found that mRNA expression of CDK2 and PCNA in LCs was increased remarkably by pcDNA3.1(+)-IGF1 treatment, whereas their transcripts were significantly weakened by si-IGF1 treatment, implicating that IGF1 can shorten the cell cycle and promote the proliferation of LCs by regulating the expression of CDK2 and PCNA.
IGF1R, also known as type 1 receptor, is a tyrosine kinase that is expressed ubiquitously on the PI3K-AKT and ERK signaling pathway. Research has indicated that IGF1 and IGF1R proteins are expressed in spermatogonia of the mature testis, and may promote spermatogenesis and regulate Leydig cell function in adult dogs37. Deficiency in IGF1R is associated with developmental delay and mental decline, while its overactivation is linked to oncogenesis38,39. The results of the present study demonstrated a significant increase in the mRNA and protein expression levels of IGF1R following the overexpression of IGF1. Conversely, a decrease in IGF1R expression was observed after silencing IGF1. These findings suggest that IGF1 may bind to the pathway receptors IGF1R to activate the PI3K/AKT and ERK signaling pathway, which in turn regulates cell growth and spermatogenesis.
In males, the decrease in serum testosterone levels with age is a complex process, partly due to a significant decrease in the ability of aging Leydig cells to produce testosterone. Many studies have shown that testosterone deficiency can cause male hypogonadism and aging40. Recent publications studies have informed us that in middle-aged and older men with decline testosterone levels revealed efficacy therapy modest improvement sexual function41. The study found that the number of spermatogonia in the testes of tilapiaOreochromis niloticus gradually increased with the increase of testosterone levels42. Study has demonstrated that cytochrome P450 family 11 subfamily A member 1 gene (CYP11A1) can metabolize cholesterol in the mitochondrial inner membrane into pregnenolone, which increases the precursor for testosterone synthesis43. In human adrenocortical carcinoma cells, Fluorene-9-bispyrone (BHPF) inhibits testosterone production by down-regulating CYP11A1 gene expression44. Steroid Reductase Domain Containing Protein 5 A (SRD5A), which plays a role in catalyzing the reduction of steroid hormones using NADPH as a cofactor45. Studies have shown that the combination of eucommia ulmoides and achyranthes japonica can enhance testosterone synthesis in aged rats by reducing the expression level of SRD5A2 gene46. Since the expression of SRD5A2 and CYP11A1 was significantly down-regulated and up-regulated in the overexpression group, resulting in differences in the expression of testosterone synthesis-related genes and testosterone content between the overexpression and control groups. Considered together, the findings from peers and us indicate that IGF1 may affect the content of testosterone in LCs by regulating the expression of SRD5A2 and CYP11A1.
Transcriptomics (RNA-seq) analysis was performed to explore the function of IGF1 in mammalian testis Leydig cells. The result showed that 1,040 DEGs were identified in the pcDNA3.1(+)-and LCs reoups, which were associated with 358 significant GO terms and 40 KEGG pathways. Some terms and pathways mainly associated with biological processes like developmental process, tight junction, extracellular matrix components, and NADPH oxidation. Specifically, the signaling pathways related to aldosterone synthesis and secretion, PI3K-Akt signaling pathway, ABC transporters, and Metabolic pathways showed significant enrichment of DEGs. There are numerous junctional molecules involved in forming the interactions and communication between germ and somatic cells, facilitate the migration of developing germ cells from the basal to the abluminal compartment of the seminiferous epithelium47. Extracellular matrix (ECM) serves as a structural support and dynamic microenvironment for cells, regulating their behavior and fate48. It plays a crucial role in various fundamental processes, including its function as a critical component of the stem cell niche, including the ECM maintains stem cells and triggers their proliferation and differentiation in response to specific stimuli49. The PI3K-Akt signaling pathway can transmit extracellular stimuli through the cell membrane to the cytoplasm, and plays an important role in testicular development and spermatogenesis50. Collectively, our results implied that DEGs were participated in the regulation of LCs development and testosterone synthesis through a variety of pathways.
To get a more detailed insight into the significance of the identified DEGs and mine the functional candidate genes implicated in testosterone synthesis, we constructed a PPI network based on their assigned pathways and functional modules. Among these, 181 testosterone synthesis-related genes were used to establish the PPI network by using Cytoscape and five genes in the PPI network were identified as hub genes, including PRKCA, CALM3, CAMK2D, PRKACA, and PRKACB. It has been shown that PRKCA (protein kinase Cα), a protein kinase widely found in cells, is an essential regulator for maintaining the integrity of the reproductive system and secondary sexual characteristics in male animals, and plays an important role in the conversion of cholesterol to testosterone. Studies have shown that arsenic can affect testosterone synthesis in mice by reducing the phosphorylation of protein kinase Cα, thereby interfering with spermatogenesis51. Previous study demonstrated that calmodulin is involved in regulating the transport of cholesterol to mitochondria in rat Leydig cells52. This network also contained the genes enriched in cellular oxidative stress molecules (such as GSTA1 and GSTA4), Adherens junction (i.e., PRKACA, WASF3 and COL1A1), steroid hormone biosynthesis (i.e., CYP11A1, HSD11B2), cytochrome P450 (i.e., GSTA1, CYP17A1 and CYP3A4) and GnRH secretion (such as GPER1 and CACNA1H). One important reason for male infertility is oxidative stress and its destructive effects on sperm structures and functions. Under oxidative stress, cells increase the expression of GSTA1 and GSTA4, helping to scavenge free radicals and other harmful oxidants, thereby protecting testicular tissue from damage, and increasing the cell viability53. Our results indicated that PRKCA and CALM3 are upregulated in the overexpression group and hence suggestive the potential importance of IGF1 in the process of cholesterol conversion to testosterone. Given the importance of testosterone synthesis by LCs, we speculate that these identified cell oxidative stress molecules and junctions and other related genes may be involved in the regulation of interstitial cell proliferation and testosterone synthesis through changes in expression, providing a basis for the development of the reproductive system and spermatogenesis of Tibetan sheep.
This is the first report regarding molecular cloning and characteristics of the ovine IGF1 CDS region, as well as potential roles during Tibetan sheep testosterone synthesis. The full-length CDS of the Tibetan sheep IGF1 gene was 502-bp-long and encoded 154 amino acids. Transcriptome analysis revealed IGF1 stimulated the expression of the proliferating cell nuclear antigen and IGF1R and promoted the proliferation of LCs via the PI3K/Akt signaling pathway. In addition, overexpression of IGF1 increased the proliferation rate, decreased apoptosis of LCs, altered expression of genes related to testosterone synthesis, and significantly increased amount of the final product testosterone. Based on these findings, we concluded that IGF1 might play roles in the proliferation or differentiation and testosterone synthesis of Leydig cells.
The datasets generated or analyzed during the current study have been deposited in the Genome Sequence Archive of the China National Center for Bio information, accession number is CRA014833 (https://ngdc.cncb.ac.cn/gsa/search? searchTerm=+CRA014833).
Smołucha, G., Piórkowska, K., Ropka-Molik, K. & Sikora, J. Use of the HRM Method in Quick Identification of FecX(O) Mutation in Highly Prolific Olkuska Sheep. Animals(basel) 10, 844 (2020).
Fu, X., Yang, Y., Yan, Z., Liu, M. & Wang, X. Transcriptomic Study of Spermatogenesis in the Testis of Hu Sheep and Tibetan Sheep. Genes (Basel). 13, 2212 (2022).
Article PubMed CAS Google Scholar
Li, T. et al. Gene expression patterns and protein cellular localization suggest a novel role for DAZL in developing tibetan sheep testes. Gene. 731, 144335 (2020).
Article PubMed CAS Google Scholar
Khawar, M. B. et al. Sirt1 regulates testosterone biosynthesis in Leydig cells via modulating autophagy. Protein Cell. 12, 67–75 (2021).
Article PubMed CAS Google Scholar
Heinrich, A. & DeFalco, T. Essential roles of interstitial cells in testicular development and function. Andrology. 8, 903–914 (2020).
Article PubMed CAS Google Scholar
Ge, R. S. & Hardy, M. P. Variation in the end products of androgen biosynthesis and metabolism during postnatal differentiation of rat leydig cells. Endocrinology. 139, 3787–3795 (1998).
Article PubMed CAS Google Scholar
Walker, W. H. Non-classical actions of testosterone and spermatogenesis. Philos. Trans. R Soc. Lond. B Biol. Sci. 365, 1557–1569 (2010).
Article PubMed PubMed Central CAS Google Scholar
Rey, R. A. The role of Androgen Signaling in male sexual development at Puberty. Endocrinology. 162, bqaa215 (2021).
Article PubMed Google Scholar
Cannarella, R., Condorelli, R. A., La, V. & Calogero, A. E. Effects of the insulin-like growth factor system on testicular differentiation and function: a review of the literature. Andrology. 6, 3–9 (2018).
Article PubMed CAS Google Scholar
Li, T. et al. Unraveling stage-dependent expression patterns of circular RNAs and their related ceRNA modulation in ovine postnatal Testis Development. Front. Cell. Dev. Biol. 9, 627439 (2021).
Article PubMed PubMed Central Google Scholar
Zhang, Y., Tang, S., Yang, W. & Du, F. let-7b-5p suppresses the proliferation and migration of pulmonary artery smooth muscle cells via down-regulating IGF1. Clin. (Sao Paulo). 77, 100051 (2022).
Article Google Scholar
Shi, J. W. et al. An IGF1-expressing endometrial stromal cell population is associated with human decidualization. BMC Biol. 20, 276 (2022).
Article PubMed PubMed Central CAS Google Scholar
Kordowitzki, P., Krajnik, K., Skowronska, A. & Skowronski, M. T. Pleiotropic effects of IGF1 on the oocyte. Cells. 11, 1610 (2022).
Article PubMed PubMed Central CAS Google Scholar
Bahadorani, M., Hosseini, S. M., Abedi, P., Abbasi, H. & Nasr-Esfahani, M. H. Glial cell line-derived neurotrophic factor in combination with insulin-like growth factor 1 and basic fibroblast growth factor promote in vitro culture of goat spermatogonial stem cells. Growth Factors. 33, 181–191 (2015).
Article PubMed CAS Google Scholar
Zhao, X. et al. Isolation and in vitro expansion of porcine spermatogonial stem cells. Reprod. Domest. Anim. 57, 210–220 (2022).
Article PubMed CAS Google Scholar
Neirijnck, Y. et al. Tumor suppressor PTEN regulates negatively sertoli cell proliferation, testis size, and sperm production in vivo. Endocrinology. 160, 387–398 (2019).
Article PubMed CAS Google Scholar
An, X. et al. Mir-1285-3p targets TPI1 to regulate the glycolysis metabolism signaling pathway of tibetan sheep sertoli cells. PLoS ONE. 17, e0270364 (2022).
Article PubMed PubMed Central CAS Google Scholar
Yang, J. M. et al. Cadmium-induced damage to primary cultures of rat leydig cells. Reprod. Toxicol. 17, 553–560 (2003).
Article PubMed CAS Google Scholar
Michaelidou, K., Tzovaras, A., Missitzis, I., Ardavanis, A. & Scorilas, A. The expression of the CEACAM19 gene, a novel member of the CEA family, is associated with breast cancer progression. Int. J. Oncol. 42, 1770–1777 (2013).
Article PubMed CAS Google Scholar
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods. 12, 357–360 (2015).
Article PubMed PubMed Central CAS Google Scholar
Anders, S., Pyl, P. T. & Huber, W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics. 31, 166–169 (2015).
Article PubMed CAS Google Scholar
Love, M. I., Huber, W. & Anders, S. Moderated estimation of Fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Article PubMed PubMed Central Google Scholar
Kanehisa, M. et al. KEGG for linking genomes to life and the environment. Nucleic Acids Res. Jan. 36, D480–D484 (2008).
Article CAS Google Scholar
Neirijnck, Y., Papaioannou, M. D. & Nef, S. The Insulin/IGF system in mammalian sexual Development and Reproduction. Int. J. Mol. Sci. 20, 4440 (2019).
Article PubMed PubMed Central CAS Google Scholar
Hu, G. X. et al. Deletion of the Igf1 gene: suppressive effects on adult Leydig cell development. J. Androl. 31, 379–387 (2010).
Article PubMed PubMed Central CAS Google Scholar
Yu, H., Zhao, Y., Guo, C., Gan, Y. & Huang, H. The role of proline substitutions within flexible regions on thermostability of luciferase. Biochim. Biophys. Acta. 1854, 65–72 (2015).
Article PubMed CAS Google Scholar
Print, C. G. & Loveland, K. L. Germ cell suicide: new insights into apoptosis during spermatogenesis. Bioessays. 22, 423–430 (2000).
Article PubMed CAS Google Scholar
Ross, A. J. et al. BCLW mediates survival of postmitotic sertoli cells by regulating BAX activity. Dev. Biol.239, 295–308 (2001).
Article PubMed CAS Google Scholar
Knudson, C. M., Tung, K. S., Tourtellotte, W. G., Brown, G. A. & Korsmeyer, S. J. Bax-deficient mice with lymphoid hyperplasia and male germ cell death. Science. 270, 96–99 (1995).
Article ADS PubMed CAS Google Scholar
Li, T. Identification and expression regulation of testicular functional genes in tibetan sheep based on transcriptomics. [D]. Gansu Agricultural University. 2021.
Johnson, C., Kastelic, J. & Thundathil, J. Role of Akt and mammalian target of rapamycin signalling in insulin-like growth factor 1-mediated cell proliferation in porcine sertoli cells. Reprod. Fertil. Dev. 32, 929–940 (2020).
Article PubMed CAS Google Scholar
Ren, F. et al. Single-cell transcriptomics reveals male germ cells and sertoli cells developmental patterns in dairy goats. Front. Cell. Dev. Biol. 10, 944325 (2022).
Article PubMed PubMed Central Google Scholar
Ma, X., Tang, T. S. & Guo, C. Regulation of translesion DNA synthesis in mammalian cells. Environ. Mol. Mutagen. 61, 680–692 (2020).
Article PubMed CAS Google Scholar
Leng, X. et al. Mouse Tspyl5 promotes spermatogonia proliferation through enhancing Pcna-mediated DNA replication. Reprod. Fertil. Dev. 36, RD23042 (2024).
Article PubMed Google Scholar
Qiao, D. H. et al. Mir-1301-3p suppresses tumor growth by downregulating PCNA in thyroid papillary cancer. Am. J. Otolaryngol. 42, 102920 (2021).
Article PubMed CAS Google Scholar
Chohan, T. A., Qian, H., Pan, Y. & Chen, J. Z. Cyclin-dependent kinase-2 as a target for cancer therapy: progress in the development of CDK2 inhibitors as anti-cancer agents. Curr. Med. Chem. 22, 237–263 (2015).
Article PubMed CAS Google Scholar
Müller, L., Kowalewski, M. P., Reichler, I. M., Kollár, E. & Balogh, O. Different expression of leptin and IGF1 in the adult and prepubertal testis in dogs. Reprod. Domest. Anim. 52, 187–192 (2017).
Article PubMed Google Scholar
López, I. P. et al. Differential organ phenotypes after postnatal Igf1r gene conditional deletion induced by tamoxifen in UBC-CreERT2; Igf1r fl/fl double transgenic mice. Transgenic Res. 24, 279–294 (2015).
Article PubMed Google Scholar
Mora, M. et al. Aging phenotype and its relationship with IGF-I gene promoter polymorphisms in elderly people living in Catalonia. Growth Horm. IGF Res. 21, 174–180 (2011).
Article PubMed CAS Google Scholar
Gryzinski, G. M. & Bernie, H. L. Testosterone deficiency and the aging male. Int. J. Impot. Res. 34, 630–634 (2022).
Article PubMed CAS Google Scholar
Maria, G. F., Thiago, G. J. & Shehzad, B. Male Reproduction and Aging. Endocrinol. Metab. Clin. North. Am. 52, 211–228 (2023).
Article Google Scholar
Nakamura, M. & Nagahama, Y. Differentiation and development of Leydig cells, and changes of testosterone levels during testicular differentiation in tilapiaOreochromis Niloticus. Fish. Physiol. Biochem.7, 211–219 (1989).
Article PubMed CAS Google Scholar
Di, G., Xiang, J., Dong, L. & Wu, J. Testosterone synthesis in testicular leydig cells after long-term exposure to a static electric field (SEF). Toxicology. 458, 152836 (2021).
Article PubMed CAS Google Scholar
Yuan, H. et al. Fluorene-9-bisphenol regulates steroidogenic hormone synthesis in H295R cells through the AC/cAMP/PKA signaling pathway. Ecotoxicol. Environ. Saf. 243, 113982 (2022).
Article Google Scholar
Ren-Shan, G. Variation in the End products of Androgen Biosynthesis and metabolism during postnatal differentiation of Rat Leydig cells. Endocrinology. 139, 3787–3795 (1998).
Article Google Scholar
Jeong, L. et al. The Effect of a Combination of Eucommia ulmoides and Achyranthes japonica on Alleviation of Testosterone Deficiency in Aged Rat Models. Nutrients.4, 3341 (2022).
Xiang, X., Dolores, D. M. & Cheng, C. Y. Intercellular adhesion molecules (ICAMs) and spermatogenesis. Hum. Reprod. Update. 19, 167–186 (2013).
Article Google Scholar
Francesca, G., Anna, U. & Paolo, B. Extracellular matrix: a dynamic microenvironment for stem cell niche. Biochim. Biophys. Acta. 1840, 2506–2519 (2014).
Article Google Scholar
Mark, H. M. et al. Human testis Extracellular Matrix enhances human Spermatogonial Stem Cell Survival in Vitro. Tissue Eng. Part. A. 25, 663–676 (2019).
Article Google Scholar
Jung-Hsuan, C. et al. LCRMP-1 is required for spermatogenesis and stabilises spermatid F-actin organization via the PI3K-Akt pathway. Commun. Biol. 6, 389 (2023).
Article Google Scholar
Xiujuan, L. et al. Arsenic interferes with spermatogenesis involving Rictor/mTORC2-mediated blood-testis barrier disruption in mice. Ecotoxicol. Environ. Saf. 257, 114914 (2023).
Article Google Scholar
Peter, F. H., Shôji, O. & James, J. M. The influence of calmodulin on steroid synthesis in leydig cells from rat testis. Endocrinology. 109, 1677–1682 (1981).
Article Google Scholar
Zhu, Z. et al. Glutamine protects rabbit spermatozoa against oxidative stress via glutathione synthesis during cryopreservation. Reprod. Fertil. Dev. 29, 2183–2194 (2017).
Article PubMed CAS Google Scholar
Download references
The research was funded by the National Natural Science Foundation of China (32260833) and Discipline Team Project of Gansu Agricultural University (GAU-XKTD-2022-20).
College of Animal Science and Technology, Gansu Agricultural University, Lanzhou, 730070, China
Yi Wu, Qiao Li, Xingcai Qi, Zilong Liu, Chunhui Wang, Xingxu Zhao & Youji Ma
Gansu Key Laboratory of Animal Generational Physiology and Reproductive Regulation, Lanzhou, 730070, China
Yi Wu, Qiao Li, Xingcai Qi, Zilong Liu, Chunhui Wang, Xingxu Zhao & Youji Ma
You can also search for this author in PubMed Google Scholar
You can also search for this author in PubMed Google Scholar
You can also search for this author in PubMed Google Scholar
You can also search for this author in PubMed Google Scholar
You can also search for this author in PubMed Google Scholar
You can also search for this author in PubMed Google Scholar
You can also search for this author in PubMed Google Scholar
“Y.W. and Y.M conceived the experiment(s), Y.W, Q.L, X.Z and C.W. conducted the experiment(s), Y.W, X.Q. and Z.L. analyzed the results, Y.W. and Y.M writing-review and editing. “.
Correspondence to Youji Ma.
The authors declare no competing interests.
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Below is the link to the electronic supplementary material.
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
Reprints and permissions
Wu, Y., Li, Q., Qi, X. et al. Molecular characteristics and regulatory role of insulin-like growth factor 1 gene in testicular Leydig cells of Tibetan sheep. Sci Rep 14, 24799 (2024). https://doi.org/10.1038/s41598-024-75234-2
Download citation
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-75234-2
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
Advertisement
© 2025 Springer Nature Limited
Sign up for the Nature Briefing newsletter — what matters in science, free to your inbox daily.

More Stories
From Commitments to Practice: Internet Society’s Priorities for WSIS+20 Implementation
Final Results of the 2026 Internet Society Board of Trustees Elections and IETF Selections
Community Snapshot—March